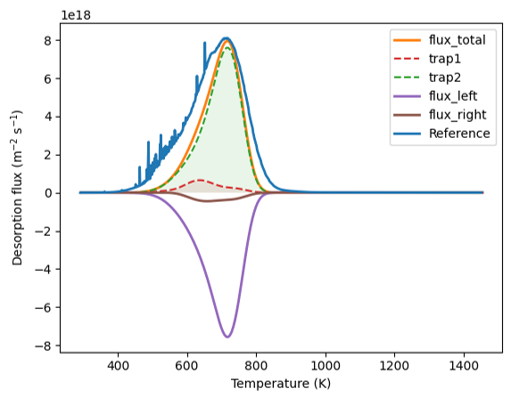
Hi everyone, I am trying to reproduce a series of plasma irradiation experiments on tungsten and TDS.(The goal is to reproduce the TDS spectrum.)
I am trying to reproduce a series of plasma irradiation and TDS with reference to Example 2 of the workshop.
However, I have some problems.
(1) When the value of E_p is reduced and the position of the peak is moved to the lower temperature side, the size of the peak should also be reduced.(I want to move the peak position without changing the peak size.)
To solve this problem, I tried changing the density of defects, but it did not work.
(2) I tried to write an auto-fitting code based on example 10 in the workshop. However, this also does not seem to work.
The experiment scenario is as follows.
The process is
\begin{equation} \begin{cases} \text{Implantation}, & \text{if} \: t < 3260 \\ \text{Cooling}, & \text{elif} \: 3260 <= t < 3295 \\ \text{Resting}, & \text{elif} \: 3295 <= t < 10460 \\ \text{TDS (1K/s)}, & \text{else} \: 10460 <= t < 11620 \\ \end{cases} \end{equation}
And the specimen temperature is
\begin{equation} T(t) = \begin{cases} 644, & \text{if} \: t < 3260 \\ 644-10(t-3260), & \text{elif} \: 3260 <= t < 3295 \\ 294, & \text{elif} \: 3295 <= t < 10460 \\ 294+(t-10460), & \text{else} \: 10460 <= t < 11620 \\ \end{cases} \end{equation}
T is expressed in \text{K}.
Implantation flux is
\begin{equation} \begin{cases} (1-r)6.1 \times 10^{21}\: \mathrm{D\:s^{-1}m^{-2}}, & \text{if} \: t < 3260 \\ 0, & \text{else}\\ \end{cases} \end{equation}
Then, r is reflectance
Implantation fluence is 2 \times 10^{25} \: \mathrm{D/m^2}.
and, Sample thickness is 0.2 \: \mathrm{mm}.
The code is shown below. Also. I also share the CSV of the reference data that is the subject of the reproduction.
TDS model
import festim as F
import numpy as np
import sympy as sp
import h_transport_materials as htm
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
def TDS(n1, E_p1, n2, E_p2):
w_atom_density = 6.3e28 # atom/m3
trap_conc1 = n1 * w_atom_density
trap_conc2 = n2 * w_atom_density
D = (htm.diffusivities.filter(material="tungsten").filter(author="frauenfelder")[0])
# Define Simulation object
synthetic_TDS = F.Simulation()
# Define a simple mesh
vertices=np.linspace(0, 0.2e-3, num=1000)
synthetic_TDS.mesh = F.MeshFromVertices(vertices)
# Define material properties
tungsten = F.Material(
id=1,
D_0=D.pre_exp.magnitude, # m2/s
E_D=D.act_energy.magnitude, # eV
)
synthetic_TDS.materials = tungsten
# Define traps
trap_1 = F.Trap(
k_0=D.pre_exp.magnitude / (1.1e-10**2 * 6 * w_atom_density),
E_k=D.act_energy.magnitude,
p_0=1e13,
E_p=E_p1,
density=trap_conc1,
materials=tungsten,
)
trap_2 = F.Trap(
k_0=D.pre_exp.magnitude / (1.1e-10**2 * 6 * w_atom_density),
E_k=D.act_energy.magnitude,
p_0=1e13,
E_p=E_p2,
density=trap_conc2,
materials=tungsten,
)
synthetic_TDS.traps = [trap_1, trap_2]
# Set initial conditions
implantation_time = 3260 # s
r = 0.5
ion_flux = sp.Piecewise(((1-r)*6.1e21, F.t <= implantation_time), (0, True))
# Set boundary conditions
synthetic_TDS.boundary_conditions = [
F.ImplantationDirichlet(surfaces=1, phi=ion_flux, R_p=4.5e-9, D_0=D.pre_exp.magnitude, E_D=D.act_energy.magnitude),
F.DirichletBC(surfaces=2, value=0, field=0)
]
# Define the material temperature evolution
implantation_temp = 644 # K
temperature_ramp = 1 # K/s
cooling_ramp = 10 # K/s
cooling_time = 350/cooling_ramp
start_tds = implantation_time + 7200 # s
synthetic_TDS.T = F.Temperature(
value=sp.Piecewise(
(implantation_temp, F.t < implantation_time),
(implantation_temp - cooling_ramp*(F.t-implantation_time), (implantation_time <= F.t)&(F.t < implantation_time+cooling_time)),
(implantation_temp - cooling_ramp*(implantation_time+cooling_time-implantation_time), (implantation_time+cooling_time <= F.t)&(F.t < start_tds)),
(implantation_temp - cooling_ramp*(implantation_time+cooling_time-implantation_time) + temperature_ramp*(F.t-start_tds), True),
)
)
# Define the simulation settings
synthetic_TDS.dt = F.Stepsize(
initial_value=0.01,
stepsize_change_ratio=1.2,
max_stepsize=lambda t: 5 if t >= start_tds else None,
dt_min=1e-08,
milestones=[implantation_time, start_tds],
)
synthetic_TDS.settings = F.Settings(
absolute_tolerance=1e11,
relative_tolerance=1e-10,
final_time=11620,
maximum_iterations=50,
)
# Define the exports
derived_quantities = F.DerivedQuantities(
[
F.TotalVolume("solute", volume=1),
F.TotalVolume("retention", volume=1),
F.TotalVolume("1", volume=1),
F.TotalVolume("2", volume=1),
F.HydrogenFlux(surface=1),
F.HydrogenFlux(surface=2),
F.AverageVolume(field="T", volume=1)
]
)
synthetic_TDS.exports = [derived_quantities]
synthetic_TDS.initialise()
synthetic_TDS.run()
return derived_quantities
Generate dummy data
# Get the flux dependence
reference_prms = [8.0e-4, 0.85, 5.0e-4, 1.2]
data = TDS(*reference_prms)
Plot dummy data
import matplotlib.pyplot as plt
import os
# Get temperature
T = data.filter(fields="T").data
# Calculate the total desorptio flux
flux_left = data.filter(fields="solute", surfaces=1).data
flux_right = data.filter(fields="solute", surfaces=2).data
flux_total = -(np.array(flux_left) + np.array(flux_right))
t = data.t
implantation_time = 3260 # s
start_tds = implantation_time + 7200 # s
tds_start_ind = t.index(start_tds)
T = T[tds_start_ind:]
flux_left = flux_left[tds_start_ind:]
flux_right = flux_right[tds_start_ind:]
flux_total = flux_total[tds_start_ind:]
trap_1 = data.filter(fields="1").data
trap_2 = data.filter(fields="2").data
contribution_trap_1 = -np.diff(trap_1)/np.diff(t)
contribution_trap_2 = -np.diff(trap_2)/np.diff(t)
contribution_trap_1 = contribution_trap_1[tds_start_ind-1:]
contribution_trap_2 = contribution_trap_2[tds_start_ind-1:]
# Visualise
plt.plot(T, flux_total, linewidth=2, label='flux_total')
plt.plot(T, contribution_trap_1, linestyle="--", color="red", label='trap1')
plt.fill_between(T, 0, contribution_trap_1, facecolor='red', alpha=0.1)
plt.plot(T, contribution_trap_2, linestyle="--", color="orange", label='trap2')
plt.fill_between(T, 0, contribution_trap_2, facecolor='orange', alpha=0.1)
plt.legend()
plt.ylabel(r"Desorption flux (m$^{-2}$ s$^{-1}$)")
plt.xlabel(r"Temperature (K)")
plt.show()
Automated TDS fit
ref = np.genfromtxt("tds-ref-fit.csv", delimiter=";")
ref[0][0]=ref[1][0] # ref[0][0]=nan without this script.
def info(i, p):
"""
Print information during the fitting procedure
"""
print("-" * 40)
print(f"i = {i}")
print("New simulation.")
print(f"Point is: {p}")
from scipy.interpolate import interp1d
prms = []
errors = []
def error_function(prm):
"""
Compute average absolute error between simulation and reference
"""
global i
global prms
global errors
prms.append(prm)
i += 1
info(i, prm)
# Filter the results if a negative value is found
if any([e < 0 for e in prm]):
return 1e30
# Get the simulation result
n1, Ep1, n2, Ep2 = prm
res = TDS(n1, Ep1, n2, Ep2)
implantation_time = 3260 # s
start_tds = implantation_time + 7200 # s
tds_start_ind = t.index(start_tds)
T = np.array(res.filter(fields="T").data)
T = T[tds_start_ind:]
flux = -np.array(res.filter(fields="solute", surfaces=1).data) - np.array(
res.filter(fields="solute", surfaces=2).data
)
flux = flux[tds_start_ind:]
# Plot the intermediate TDS spectra
if i == 1:
plt.plot(T, flux, color="tab:red", lw=2, label="Initial guess")
else:
plt.plot(T, flux, color="tab:grey", lw=0.5)
interp_tds = interp1d(T, flux, fill_value="extrapolate")
# Compute the mean absolute error between sim and ref
err = np.abs(interp_tds(ref[:, 0]) - ref[:, 1]).mean()
print(f"Average absolute error is : {err:.2e}")
errors.append(err)
return err
from scipy.optimize import minimize
i = 0 # initialise counter
# Set the tolerances
fatol = 1e18
xatol = 1e-3
initial_guess = [8.0e-4, 0.85, 5.0e-4, 1.2]
# Minimise the error function
res = minimize(
error_function,
np.array(initial_guess),
method="Nelder-Mead",
options={"disp": True, "fatol": fatol, "xatol": xatol},
)
# Process the obtained results
predicted_data = TDS(*res.x)
T = predicted_data.filter(fields="T").data
T = T[tds_start_ind:]
flux_left = predicted_data.filter(fields="solute", surfaces=1).data
flux_right = predicted_data.filter(fields="solute", surfaces=2).data
flux_total = -(np.array(flux_left) + np.array(flux_right))
flux_total = flux_total[tds_start_ind:]
# Visualise
plt.plot(ref[:, 0], ref[:, 1], linewidth=2, label="Reference")
plt.plot(T, flux_total, linewidth=2, label="Optimised")
plt.ylabel(r"Desorption flux (m$^{-2}$ s$^{-1}$)")
plt.xlabel(r"Temperature (K)")
plt.legend()
plt.show()
Thank you very much if someone has an idea.